Cleanrooms, DNA
How Cleanrooms are Revolutionizing DNA Analysis
Against a neutral blue background, the young man in a pressed, button-down shirt and polka dotted tie smiles the hesitant smile of the new hire. Blue eyes matching his suit blazer and blond hair slicked to one side, Scott Johnson’s fresh-faced, preppy college look belied the strength of his ambition and his drive to succeed. A passionate world traveler and avid sports fan, Johnson had reason to smile. Having secured a much-vaunted position at Keefe, Bruyette & Woods, a boutique investment bank and broker-dealer to the financial services sector, he had set up his office in Manhattan’s World Trade Center and the future was looking bright. However, the dreams of the 26-year old securities analyst were also reduced to rubble on the day the towers fell, a wallet and his employer’s HR headshot being all that remained. One of 67 Keefe, Bruyette & Woods’ employees to perish that day, Johnson’s body was never found and his family continued to struggle with finding closure. But according to a recently published article, new techniques pioneered by New York City’s Chief Medical Examiner Barbara Sampson and the Department of Forensic Biology, have finally tied a bone fragment to Johnson’s genetic profile by using new digestion chemicals to isolate the DNA from the fragment.(1) Seventeen years after the attack, Johnson’s remains have finally been identified, and some degree closure has been brought to his still grieving family. And this is not the only new development in the field of genetics and genome sequencing made possible by new techniques in DNA extraction. Let’s take a broader look…
Anyone familiar with the fields of science fiction, biopunk, or eugenics will have stumbled across the work of Andrew Niccol, the writer and director of the often-overlooked film, Gattaca. Released in 1997, Gattaca examines the tension between genetics as destiny and the role of science within a society where heredity is king, and derives its name from the leading characters of the four nucleobases of DNA: guanine (G), thymine (T), adenine (A), and cytosine (C). The order – or sequence – of these chemical bases determines the information which is available to build the cells that in turn create every living thing. The four bases join together – A and T, C and G – to form base pairs that attach to a sugar and a phosphate molecule to create a nucleotide. In turn, these nucleotides are arranged into two spiraling strands, graphically rendered as a ladder, which we term the double helix. In this ladder representation, the rungs are formed by the base pairs with the vertical sides being composed of the sugar and phosphate molecules. DNA is found in almost every cell in the human body – either in the nucleus (‘nuclear DNA’) or in the mitochondria (‘mtDNA’) – and as such is an ever-present marker of an individual life.
So, with the significance of DNA in mind, recent advances in evolutionary genetics – especially in terms of DNA extraction and analysis – are especially interesting. And they are universally applicable – whether the analysis stretches as far back in time as examining the links between Neanderthal genetics and those of modern man, the screening of human DNA in sediments where even bones are absent, or more contemporary uses such as identifying bone fragments from the Twin Towers. How? Let’s reach far back in time and take a look…
In a lab at the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, Professor Svante Pääbo is leading a team intent upon understanding why humans as a species have been so successful in colonization and the creation of cultures and societies.
Examining the differences in three genes associated with brain development, Pääbo’s team is using Crispr to edit human stem cells by changing a single letter pair such that they more closely resemble those of the Neanderthal. Bathed in a protein-rich medium, the stem cells become neurons which ‘self-organise into miniature brain-like structures that grow to a few millimeters in diameter,’ which the team terms ‘organoids.’(2) After nine months in a growing medium, a comparison can be made of the human organoids and those of the edited Neanderthal-like organoids. Studying the synapses, electrical activity, and the ways in which the cells divide, develop, and organize helps researchers understand how the two brains would have become differently wired, allowing us to infer ways in which these differences would have influenced both subjects’ ability to thrive.
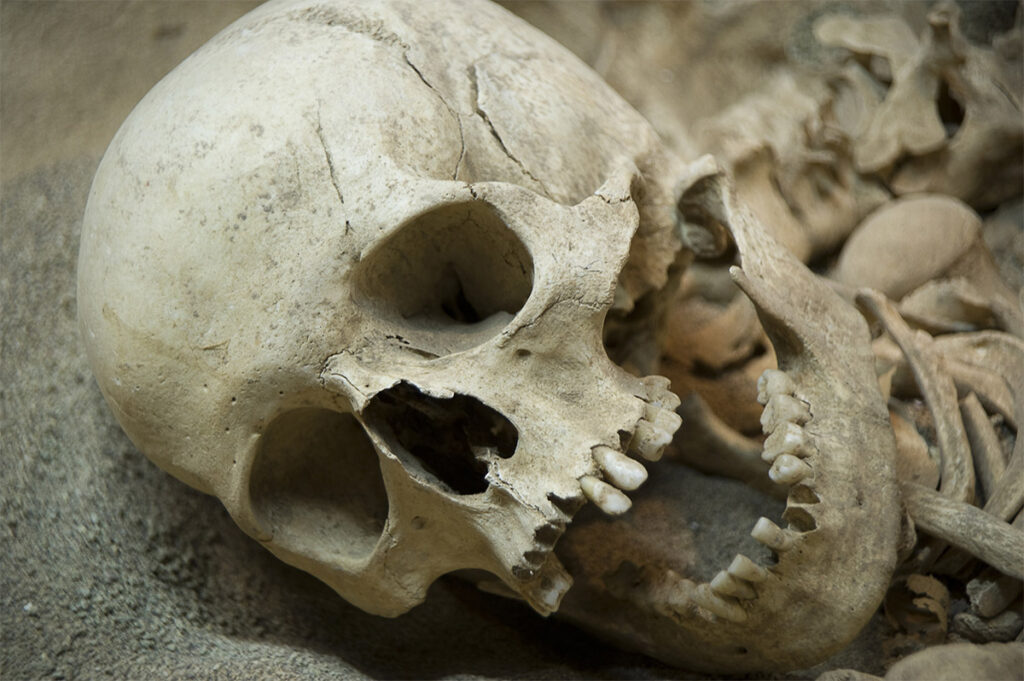
And this work is only possible in a contamination-controlled environment. As we would expect, Pääbo’s team makes full use of air showers and complete personal protective equipment (bunny suits and more) in a sterile environment, controlled by full HEPA filtration. And this is critical because, when working with the genetic material extracted from bone samples millennia old, even a microscopic particle of dust can contaminate an irreplaceable sample to the point that it is rendered useless. Protecting source materials, especially when it comes from individuals who lived and died during the mid-late Pleistocene era, is of paramount importance. After all, without the bones there can be no analysis.
Or can there?
In more groundbreaking work on the genetics of the Neanderthals and other ‘archaic’ humans such as the Denisovans, new techniques have enabled the sampling of DNA from sediments in soil samples. Picture the scene: more than 50,000 years before the advent of the port-a-potty, a wandering Neanderthal found himself in a bit of a pickle. Ducking into the relative privacy of a nearby cave, he sought relief from his discomfort and continued on his way utterly unaware that modern day humans would find and sequence his DNA. How? Through environmental DNA.
Until recently there has been no surefire way to isolate ancient DNA from accidental contamination by modern DNA that can occur during collection. However, in new work by Max Planck researchers Matthias Meyer and Viviane Slon, a ‘DNA hook’ is crafted from modern human genetic material and used to isolate sequences to which it is most similar. These are then compared against known mitochondrial DNA sequences from the target populations. And, as we’ll touch on shortly, the use of mtDNA is effective because it is much more abundant than its nuclear counterpart. And, as Hendrik Poinar of McMaster University in Hamilton, Canada, notes in an article by Lizzie Wade in Science Magazine, this shift in focus opens up myriad exciting new avenues in genetic archeology. ‘[A]ncient DNA from sediments will help [researchers] complete the map of ancient human occupations and allow them to see where species may have overlapped and interacted.’(3) Previously, the use of bones for analysis was inherently limiting insofar as they were site-specific and therefore confined that individual to one geographical locale. But, as we know, early humans traveled and spread their DNA widely as they populated the globe. By examining sedimentary DNA, we can not only track the movements of the archaic humans but also have greater confidence in the timelines of their lives and ultimately of their demise.
Maybe this is a good moment to skip forward in time. When you think of archeology, what’s the image that comes most frequently to mind? Ancient Egypt perhaps? But if you thought we knew everything it was possible to know about the Egyptians and their mummies it’s time to re-examine that belief. In what is thought to be the first successful extraction of its kind, an international team based in the bucolic Black Forest town of Tübingen and at the Max Planck Institute for the Science of Human History in Jena, Germany, has recovered nuclear data from three mummies found at Abusir el-Meleq, an archeological site along the Nile River south of Cairo. This development is significant because geneticists have traditionally been skeptical as to the reliability of nuclear DNA from mummies given the environmental conditions under which it is sampled. Hot and humid, the tombs are also laced with chemicals used in the embalming and mummification processes – none of which provides a space in which DNA can flourish. And yet it is this core nuclear DNA that is so prized, representing, as it does, the complete human genome. Where mitochondrial DNA is inherited only from the maternal line, nuclear DNA is the full package.
So using a dedicated contamination-controlled environment, nuclear DNA was successfully extracted from samples of bone, teeth, and soft tissues that had been irradiated with ultra violet light to prevent contamination.
Surface samples were removed for analysis and sufficient DNA was extracted to allow the team to study the ‘genetic differentiation and population changes and continuity over a 1,300-year time span before comparing the results to modern populations.’(4) And this opened up the field for researchers to chart the extent to which ancient populations were affected on the genetic level by foreign domination – for instance, by the conquest by Alexander the Great – or by engaging in trade, for example the trans-Saharan slave trade that emerged around 1300 years ago. In effect, the tiny sampling of nuclear DNA recovered from ancient macerated teeth now offers us fresh insights into our understanding of the book of human history, as it has been written so far.
Let’s fast-forward again. Approximately 800 years ago a young woman, aged between 19 and 24, died in Norway. In an article published in Laboratory Equipment, a publication ‘covering a broad array of news, products, and technologies for lab professionals,’ an artist’s rendering of the woman reveals a stunning image.(5) Olive-skinned with blue eyes and dark hair, the woman has almond-shaped eyes set above a broad nose, and an expression that is a curious mixture of dignity and challenge. When her remains were exhumed from a graveyard in Trondheim, it was determined that she succumbed to septicemia and enteric fever as a result of Salmonella Paratyphi C infection. But the interesting aspect of this case is that this disease is considered to be a danger only in the tropics of South Asia, Africa, and East Asia. So how did she come to die in this way?
In a cleanroom environment at the University of Copenhagen’s Natural History Museum, autosomal DNA was extracted from the internal pulp of two of her molars, along with some dental plaque, and a part of her femur.
After treating with NEBNext DNA preparation reagents, the team used the Illumina HiSeq 2500 and 400 platforms on multiple sequencing libraries, and with the help of EnteroBase, an online bioscience database at the University of Warwick Medical School, the woman’s DNA and her bacterium were compared against strains of Para C bacteria. Interestingly, it was found that the strain of the bacterium that killed her was ‘substantially similar’ to one currently making the rounds in our modern world and may have been triggered by jumping from an animal host to a human.(6)
So, what is the thread that ties all of this intriguing research together? From the Neanderthal sedimentary DNA to the 123h century Scandinavian Salmonella case to closure for another 911 family, the common thread is the critical role played by a controlled environment during the forensic analysis of remains. And although the facilities in each case are vastly different, they all have one thing in common: excellence. And what does this excellence look like? The facilities the University of Utah’s College of Social and Behavioral Science give us a clue.
In a dedicated Ancient DNA Cleanroom Laboratory students and staff enjoy a state of the art facility that incorporates an ISO Class 7 gowning area along with two ISO Class 6 work spaces. Within the controlled environment we see dedicated ISO Class 5 laminar flow hoods and the entire area leverages HEPA filtration to provide positive pressure airflow from ceiling-mounted filter points. Researchers conform to SOPs and best practices in moving slowly through the unidirectional airflow which proceeds from the cleanest space, through the gowning area, and finally to the outside of the controlled environment. Without exception, all personnel are provided with full PPE which are sanitized between uses using chemical bleaching agents, and the general space is ‘scrubbed’ with ultra-violet lighting for surface and air sterilization. The water is fed through a NANOPure Diamond Water System and reagents are provided as aliquots.
All of these components come together to form a facility that allows for pioneering work on the understanding of population history – including behavior and migration – through the analysis of genetic variation. Arguably, at a moment in history when our origins and genetic make-up once again seem to be increasingly more politically charged and socially relevant, this field of study offers a model of similarity grounded firmly in science. Despite some sectors’ wariness of ‘the other’, genetic/forensic archeology, with its use of hard science and advanced cleanroom technology, encourages an appreciation of our similarities and promotes the recognition that our differences are, by and large, purely skin-deep.
Are you as interested by Neanderthal DNA as we are? Who could have guessed how far archaic humans and modern humans would overlap? We’d love to hear your thoughts…
References:
- http://www.nydailynews.com/new-york/ny-metro-911-victim-identified-20180725-story.html
- https://www.theguardian.com/science/2018/may/11/scientists-to-grow-mini-brains-using-neanderthal-dna
- http://www.sciencemag.org/news/2017/04/no-bones-no-problem-dna-left-cave-soils-can-reveal-ancient-human-occupants
- https://www.wired.co.uk/article/ancient-mummy-genome
- https://www.laboratoryequipment.com/news/2018/07/young-norwegian-woman-medieval-times-had-predominantly-tropical-disease
- ibid



















HAVE AN IDEA FOR CONTENT?
We are always looking for ideas and topics to write about.
Contact Us